|
Tutorial 2: Step 3 Rename the Dataset
Default names are provided for all datasets and experiments based on either the name of the file being imported, or on the type of experiment being performed. Any item listed in the navigator can be renamed at any time. This gives you the opportunity to apply your own naming convention to the data.
Rename a Dataset
1. Right-click the Estimated: #mv < 30 | median dataset in the Experiments navigator, and select Rename Experiment from the shortcut menu. A box is drawn around the dataset name, with a blinking cursor at the end of the name.
2. Press the <Backspace> key to delete the program-generated name.
3. Type in something significant to you (e.g. 3-nearest-neighbors estimation).
4. Press <Enter> to accept this new name. The experiment is renamed with the new name.
Please note that GeneLinker™ saves all files automatically. Once an item is visible in the Experiments navigator, it has already been saved to the GeneLinker™ database. The renaming facility is for convenience. For instance, the name recommended in this example allows you to see at a glance, the type of missing value estimation which produced that dataset. This would be particularly valuable were you, for instance, comparing different methods for missing value estimation. The parameters used to generate every dataset are captured automatically by GeneLinker™ and can always be viewed by selecting the item and examining the Description Pane in the lower left of the application window.
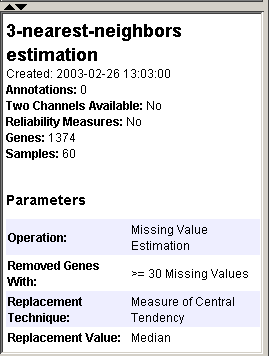
5. Do that now: The dataset is still highlighted. Look at the information provided in the Description Pane. Among other things, notice that there are 1374 genes in this dataset.
6. Click the parent dataset, ‘t_matrix’, and examine the information about it in the Description Pane. Notice that there are 1375 genes in the parent dataset.
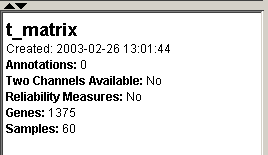
The Estimate Missing Values operation filtered out one gene because it had more missing values than we wanted. In the next step we will demonstrate one way of identifying that filtered gene.

