|
Importing Data from CodeLink XML Files
Overview
The data files must be in the CodeLink PROFILE XML file format.
CodeLink may associate up to three XML files with each slide or sample: A PATTERN file, a PROFILE file and an ID file. The PROFILE file contains the expression data which GeneLinker™ imports.
Example PROFILE.XML viewed with Microsoft Internet Explorer:

When selecting files for import, you need only select the PROFILE.XML files as in the picture below. The PATTERN and ID files should not be selected.
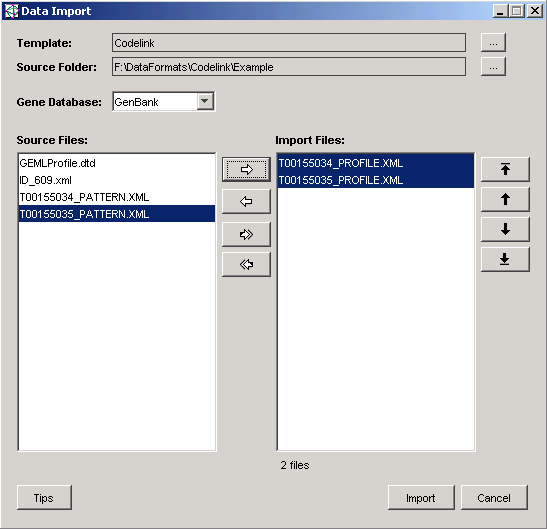
Import Process
Multiple files are processed into a single dataset. The sample order of the imported dataset is determined by the order of the source sample data files listed in the Data Import dialog as shown above.
You should use the GenBank gene database type when importing CodeLink data.
Characteristics of the CodeLink Import Template
The CodeLink import template has the following characteristics.
1. GenBank accession numbers are used as gene identifiers. These are obtained by stripping the reporter name of its '_PROBEn' extension. Although the systematic names are also GenBank accession numbers, they are sometimes non-unique: That is, two different probes may be mapped to a single systematic name. In order to preserve the distinct identities of the probes GeneLinker™ uses the reporter names. If the systematic names are desired, they can be imported as descriptions via gene list import.
2. GeneLinker™ reads the normalized iod value as the expression value. These values are already background-subtracted and normalized by division by the median value of the DISCOVERY probes on the slide.
Related Topics:
Selecting a Template for Data Import
Importing Multiple Files With One Sample Each

