|
Performing an ANOVA
Overview
This operation calculates p-values for the genes in a complete dataset. For details of the ANOVA algorithms, see Overview of ANOVA.
The input to this operation must be a complete dataset. If your dataset has missing values, see Overview of Estimating Missing Values for techniques available to eliminate or estimate missing values.
Actions
1. Click a complete dataset with variable information
identifying the replicate samples ![]() in the Experiments
navigator. The item is highlighted.
in the Experiments
navigator. The item is highlighted.
2. Select ANOVA from the Statistics menu, or right-click on the item and select ANOVA from the shortcut menu. The ANOVA dialog is displayed.
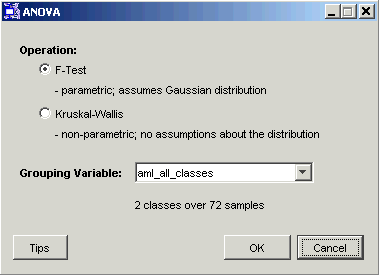
Note: if an appropriate grouping variable is not associated with the dataset, this is indicated on the dialog. In this situation, click Cancel and import an appropriate variable before trying again. See Overview of ANOVA for a discussion of appropriate variables.
3. Set the Operation (style of ANOVA) to F-Test or Kruskal-Wallis. See Overview of ANOVA for how to choose the right method.
4. Select the Grouping Variable from the drop-down list.
5. Click OK. The ANOVA operation is performed and upon successful completion, a new F-Test or Kruskal-Wallis Results item is added to the Experiments navigator under the original dataset.
The results can then be viewed using the ANOVA Viewer.
Related Topics:
Overview of Estimating Missing Values

