|
Platinum
Tutorial 6: Step 5 Display SLAM Association Viewer
If the SLAM association viewer is already displayed, there is no need to recreate it. Read the sections below the image for information about the SLAM Association Viewer.
View SLAM™ Results
1. Double-click the newly created SLAM: training classes | 30000 | 4| 0.7 item in the Experiments navigator. The item is highlighted and the SLAM association viewer is displayed.
OR
1. If the newly created SLAM: training classes | 30000 | 4| 0.7 item in the Experiments navigator is not already highlighted, click it.
2. Click the Association
Viewer toolbar icon ![]() , or select Association
Viewer from the Predict
menu, or right-click the item and select Association
Viewer from the shortcut menu. The SLAM™ association viewer is
displayed.
, or select Association
Viewer from the Predict
menu, or right-click the item and select Association
Viewer from the shortcut menu. The SLAM™ association viewer is
displayed.
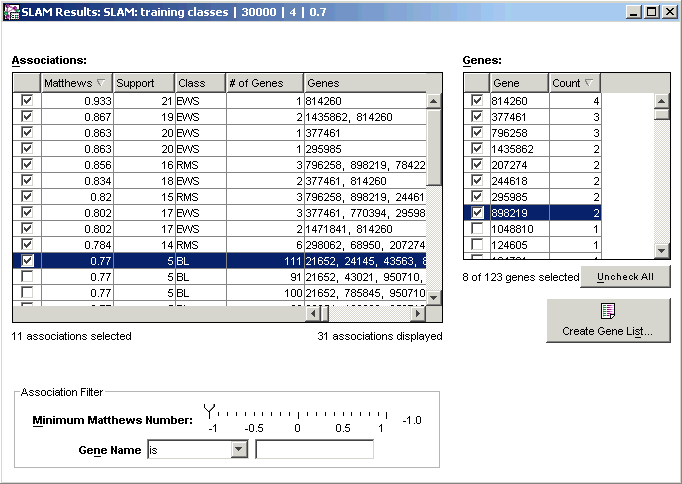
The SLAM™ Association Viewer has three functional areas:
Associations:
The Associations list displays a list of all the associations found during the SLAM™ run. To sort the list by a particular characteristic, click on the column header for that characteristic. Clicking again on the same header reverses the order of the sort (ascending or descending).
The Associations list can be sorted by:
Matthews statistic (a measure of the predictive power of the association),
Support (the number of samples in the dataset which match the pattern),
Class, or
The number of genes in the association.
Genes:
The Genes list box in the upper right lists the genes in the checked associations. A gene list can be created from the checked genes in the Genes box. The gene list can be used to identify interesting genes (features) for use in supervised learning experiments.
Note: only one copy of a gene name is listed in the Genes list box. The Count column indicates the number of associations the gene occurs within.
Association Filter
Since SLAM™ can potentially find hundreds or even thousands of associations, some methods are provided in the Association Filter group for reducing the number of associations displayed. You can display only associations with a Matthews statistic above an adjustable cutoff, or you can display only associations containing certain genes, or not containing certain genes.

