Overview
GeneLinker™ can generate two types of scripts:
A Single experiment script is a script for the experiment selected in the Experiments navigator.
For example, generating a single experiment script for a clustering experiment produces a script that includes information just about that clustering experiment.
A Workflow script is a script for the workflow leading up to and including the experiment selected in the Experiments navigator.
For example, generating a workflow script for the same clustering experiment produces a script that includes information about the original dataset, any intermediate elimination or estimation of missing values, any normalization and/or filtering steps, and the clustering experiment.
Information saved in the scripts includes (where applicable):
Experiment parameters,
Variable name,
List of genes.
Note that for scripts that use variables, such as sample merging, F-test and classifier training, the name of the variable is stored as part of the script, and to run the script on another dataset requires that the other dataset have a variable of the same name. So if you save a script that ran an F-test on a dataset with using a variable named CancerClass, to run the same script on a different dataset requires that it, too, have a variable named CancerClass.
For scripts that use genelists, for example in genelist filtering, the current list of genes defined by the genelist is saved as part of the script, not the genelist name. This means that if you delete a genelist that is used by a script, the script will continue to run as it did when it was saved. However, if you delete a genelist and create a new genelist of the same name, be aware that the script will continue to use the original list of genes, not the new one.
For scripts that involve Missing Value Estimation the fraction of missing values that triggers gene removal is saved, not the absolute number.
Scripts cannot be generated for the following experiment types:
Also, visualizations cannot be scripted, although if automatic visualization is left on when a script is run the default visualizations will be generated as part of the script's execution.
Data import and export cannot be scripted in the current version of GeneLinker™.
Actions
1. Click an item in the Experiments navigator. The item is highlighted.
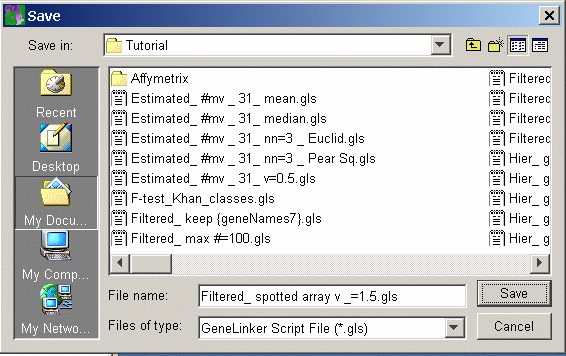
2. Select Generate Script or Generate Workflow Script from the Tools menu. The Save As dialog is displayed.

3. Navigate to the folder where the file is to be saved.
4. GeneLinker™ provides a default file name, based on the selected item’s name, with an extension of .gls. You may rename the default path and file name by typing over them.
5. Click Save. The report is saved as a GeneLinker™ Script (.gls) file in the specified folder.
Related Topics: