|
Linear Regression
Overview
This procedure scales the values across samples (gene chips) so that the slope of each sample is equivalent. This is done for all samples except the baseline.
This procedure fits a linear regression model using the intensities of the common genes in the baseline and each of the other samples. The inverse of the slope of the linear regression line becomes the (multiplicative) re-scaling factor for the current sample. The re-scaled intensity of the samples (other than baseline) becomes the original intensity multiplied by the re-scaling factor. This is done for all samples except the baseline. The baseline gets a re-scaling factor of 1.
Before clustering, it is recommended that standardization be performed after scaling using a baseline. Baseline scaling makes the intensities across chips equivalent, but genes may still differ in absolute intensity, and standardization can address this.
Actions
1. Click a complete dataset in the Experiments navigator. The item is highlighted.
2. Click the Normalize
toolbar icon ![]() , or select Normalize
from the Data menu, or right-click
the item and select Normalize
from the shortcut menu. The first Normalization
dialog is displayed.
, or select Normalize
from the Data menu, or right-click
the item and select Normalize
from the shortcut menu. The first Normalization
dialog is displayed.

3. Double-click the Sample Scaling radio button, or click it and click Next. The second Normalization dialog is displayed.
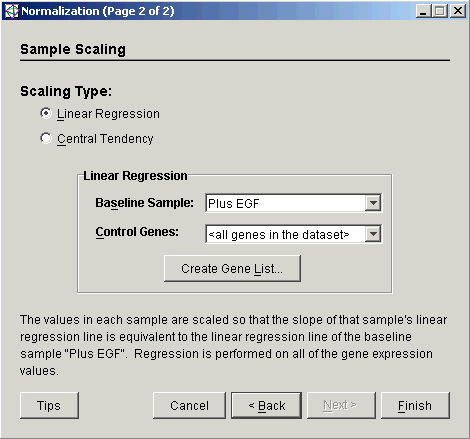
4. Select Linear Regression as the Scaling Type.
5. Set the Baseline Sample from the drop-down list. If no baseline sample is selected, the sample displayed in the box is used for the normalization operation.
6. The Control Genes (housekeeping genes) can either be 'all genes in dataset' or the genes specified in a gene list.
If 'all genes in dataset' is selected, the operation that is performed is scaling using a baseline.
If a gene list is selected, the operation that is performed is scaling using housekeeping genes. For this option, the gene list must contain at least two genes from the dataset (min. required to calculate slope) and less than all the genes in the dataset (the control genes are always discarded prior to returning the normalized dataset).
If an appropriate gene list does not exist, click Create Gene List. The Gene List Creator dialog is displayed.
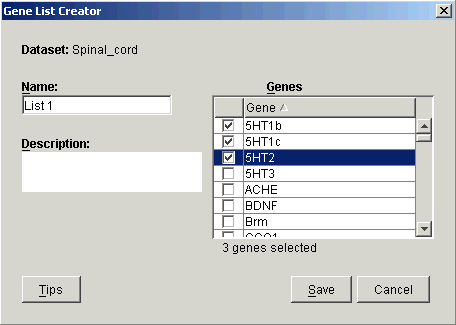
a) Type in a Name for the list and optionally a Description.
b) Click the checkboxes next to the genes to be included in the list.
c) Click Save. The gene list is then displayed in the Control Genes list on the Normalization dialog.
7. Select the Control Genes from the drop-down list.
8. Click Finish. The Experiment Progress dialog is displayed. It is dynamically updated as the Sample Scaling Normalization operation is performed. To cancel the Sample Scaling Normalization operation, click the Cancel button.
If the operation cannot complete an error message is displayed. The operation will fail, for example, if the slope of the linear regression is zero or infinity (if a sample is constant).
Upon successful completion, a new normalization dataset is added under the original dataset in the Experiments navigator.
Related Topics:

