|
Platinum
Tutorial 6: Step 12 Set URL for Lookup Gene Operation
Set URL for Lookup Gene Operation
You can create different sets of genes and evaluate the discriminant power of each by training and testing a new classifier using each gene list. You might create these alternate gene lists by running SLAM™ longer, by choosing different genes from the SLAM™ output, or from your existing knowledge of which genes participate in a given process or disease state. One way to determine what is known about a gene is to use the Lookup Gene function of GeneLinker™. If you imported your expression data using GenBank or UniGene identifiers, you can look them up simply by choosing the Lookup Gene icon. It is enabled whenever you have a gene or a gene list selected.
If you don't have GenBank or UniGene identifiers associated with your expression data, you may still be able to look up genes directly from GeneLinker™. The dataset for this tutorial, for example, uses IMAGE Consortium clone ids. Steps 12 and 13 demonstrate how GeneLinker™ can look up genes via their clone ids.
1. Select Preferences from the Tools menu. The User Preferences dialog is displayed.
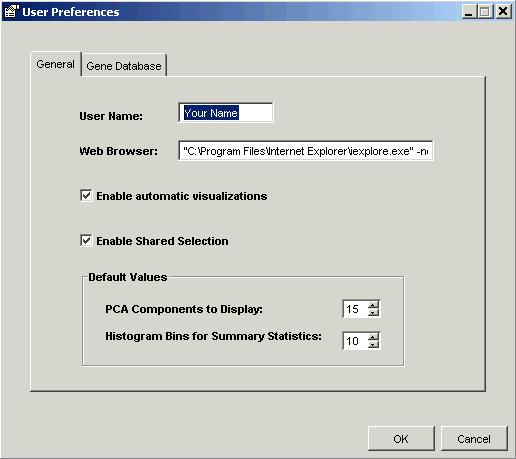
2. Click the Gene Database tab. The User Preferences dialog is updated.
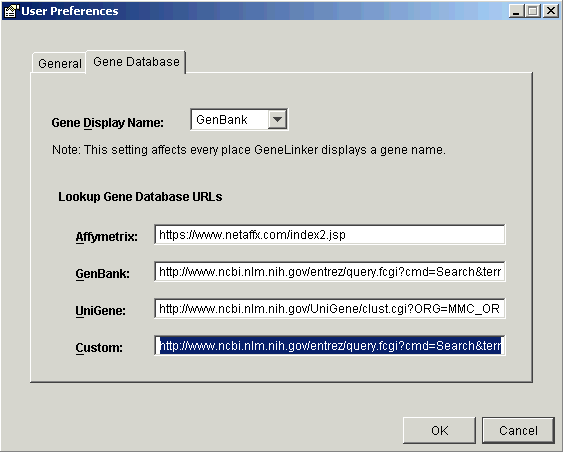
3. Under Lookup Gene Database URLs, click in the text box next to Custom. The text in the box is highlighted.
4. Either:
a) Use the right arrow key to move the cursor right until you see MMC_ID.
b) Use the mouse to highlight MMC_ID.
c) Type "IMAGE:MMC_ID" including the quotation marks.
Or:
a) Press <Delete>. The text box is cleared.
b) Copy and paste the following URL into the text box (all on a single line): http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?cmd=Search&term="IMAGE:MMC_ID"&db=Nucleotide&doptcmdl=GenBank
All that changes is that the string MMC_ID becomes "IMAGE:MMC_ID". Note the addition of the quotation marks.
Note that the URL must remain on a single line. Any line break you see in the tutorial text is due to word wrap in the GeneLinker™ Help viewer. Be sure to type the URL in on a single line.
The actual gene identifier (e.g. 207274) is substituted for the sub-string MMC_ID when you perform a Lookup Gene operation on that gene.
5. Click OK.

