|
Importing Data from Koadarray Files
Overview
The data files must be in the Koadarray file format. The Koadarray Options pane must be configured as follows in terms of the labels, Spot data types (Spot ID must be first), and filename extension:
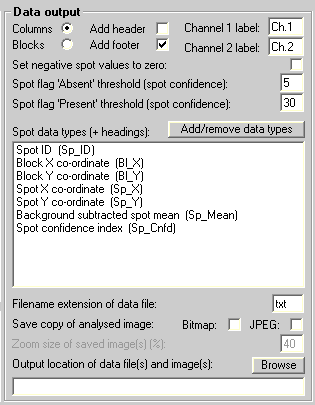
The following data file is an example of a Koadarray monochrome (one channel or ratio) data file:
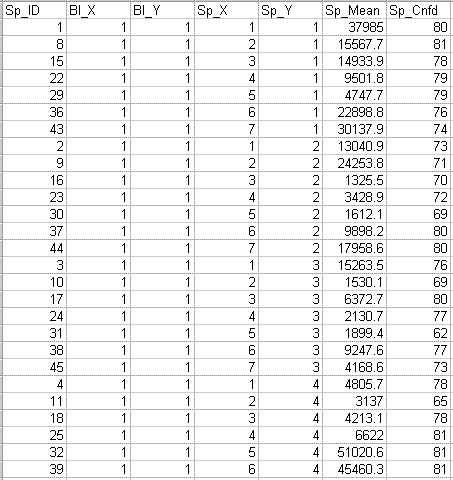
The following data file is an example of a Koadarray two color (two channel) data file:
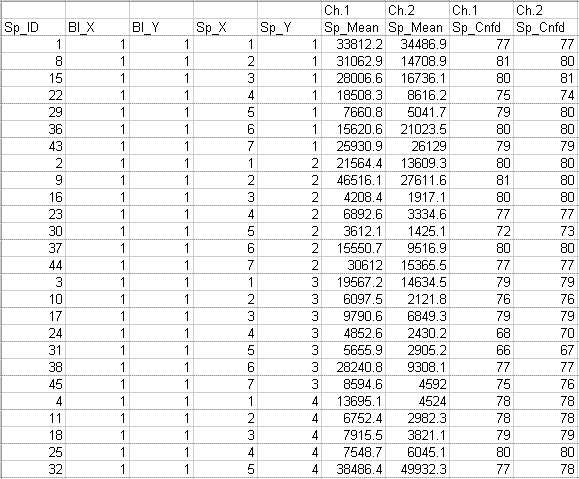
Sample Order
The sample order of imported datasets is determined by the order of the source sample data files listed in the Import Data dialog.
|
Template |
Result of Import |
|
Koadarray Ratio |
Multiple files are processed into a single dataset. |
|
Koadarray 2-Color |
Multiple files are processed into a single dataset. |
If you are importing using the two-color data template, all data values <1 (i.e. the treatment value is lower than the control) are replaced with missing values (null values). Between-chip replicate measurements are imported as samples with the same names. When the import process is complete, a dataset that is the ratio of treatment/control is added to the Experiments navigator. A selected sample ratio can be displayed in an intensity-bias plot to determine whether Lowess normalization is appropriate for the dataset.
Import Process for Koadarray Ratio Data Sets
The file headers are discarded.
Gene identifier information is retrieved from the first column (Sp_ID) of the first file and is stored as a GenBank Identifier.
Gene expression data is retrieved from the Sp_Mean column of each file in the order they are placed in the Import Data dialog.
The resulting dataset is not be amenable to Lowess Normalization or Intensity-Bias plots. See Two-Color Data for more information.
Koadarray Confidence values are normally in the form of percentages (0-100 where higher numbers represent more reliable spots). The confidence values are retrieved from the Sp_Cnfd column of each file in the order they are placed in the Import Data dialog. The Koadarray confidence values are converted as GeneLinker™ expects reliability measures to fall between 0 and 1, with 0 representing very reliable and 1 representing unreliable. This is patterned off the interpretation of p-values in traditional statistical tests, where small numbers indicate significance.
The footers are discarded if they were included in the data files.
Import Process for Koadarray 2-Color
The file headers are discarded.
Gene identifier information is retrieved from the first column (Sp_ID) of the first file and is stored as a GenBank Identifier.
The Two-Color checkbox should be checked by the user if it isn't already checked. The ratio data is calculated by dividing Ch.1Sp_Mean column by the Ch.2Sp_Mean column (or vice versa depending on the users's choice for the Two-Color Setting (Ch.1/Ch.2 or Ch.2/Ch.1). The expression data is retrieved from each file in the order that the samples are placed in the Import Data dialog.
The resulting dataset is amenable to Lowess Normalization and Intensity-Bias plots. See Two-Color Data for more information.
Koadarray Confidence values are normally in the form of percentages (0-100 where higher numbers represent more reliable spots). The confidence values are retrieved from the Ch.1Sp_Cnfd and Ch.2Sp_Cnfd columns of each file in the order they are placed in the Import Data dialog. The Koadarray confidence values are converted as GeneLinker™ expects reliability measures to fall between 0 and 1, with 0 representing very reliable and 1 representing unreliable. This is patterned off the interpretation of p-values in traditional statistical tests, where small numbers indicate significance. Since the ratio values are contructed from 2 spots (each with its own confidence values), the overall confidence value for each ratio is constructed by summing the pair of confidence values in quadrature.
The footers are discarded if they were included in the data files.
Related Topics:
Selecting a Template for Data Import
Importing Multiple Files With One Sample Each
Merging Within-Chip Replicate Measurements

